Automated Protein-Ligand MD Simulations & MM/PBSA
- 15+ Automated Trajectory Analytics
- Rigorous MM/PBSA Binding Energetics
- Publication & Audit-Ready Reports
Output
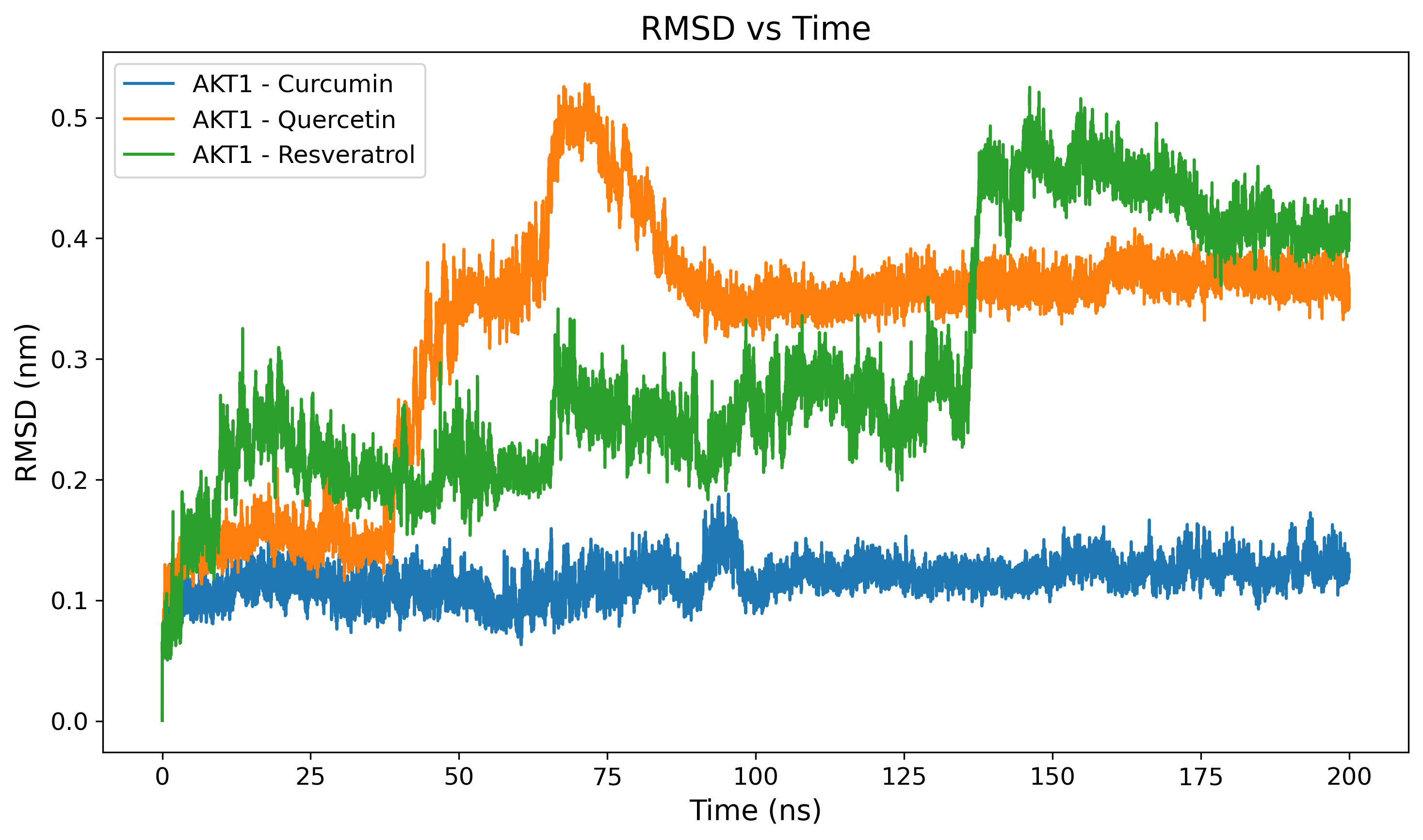
Protein RMSD
Backbone Root Mean Square Deviation to assess overall structural stability.
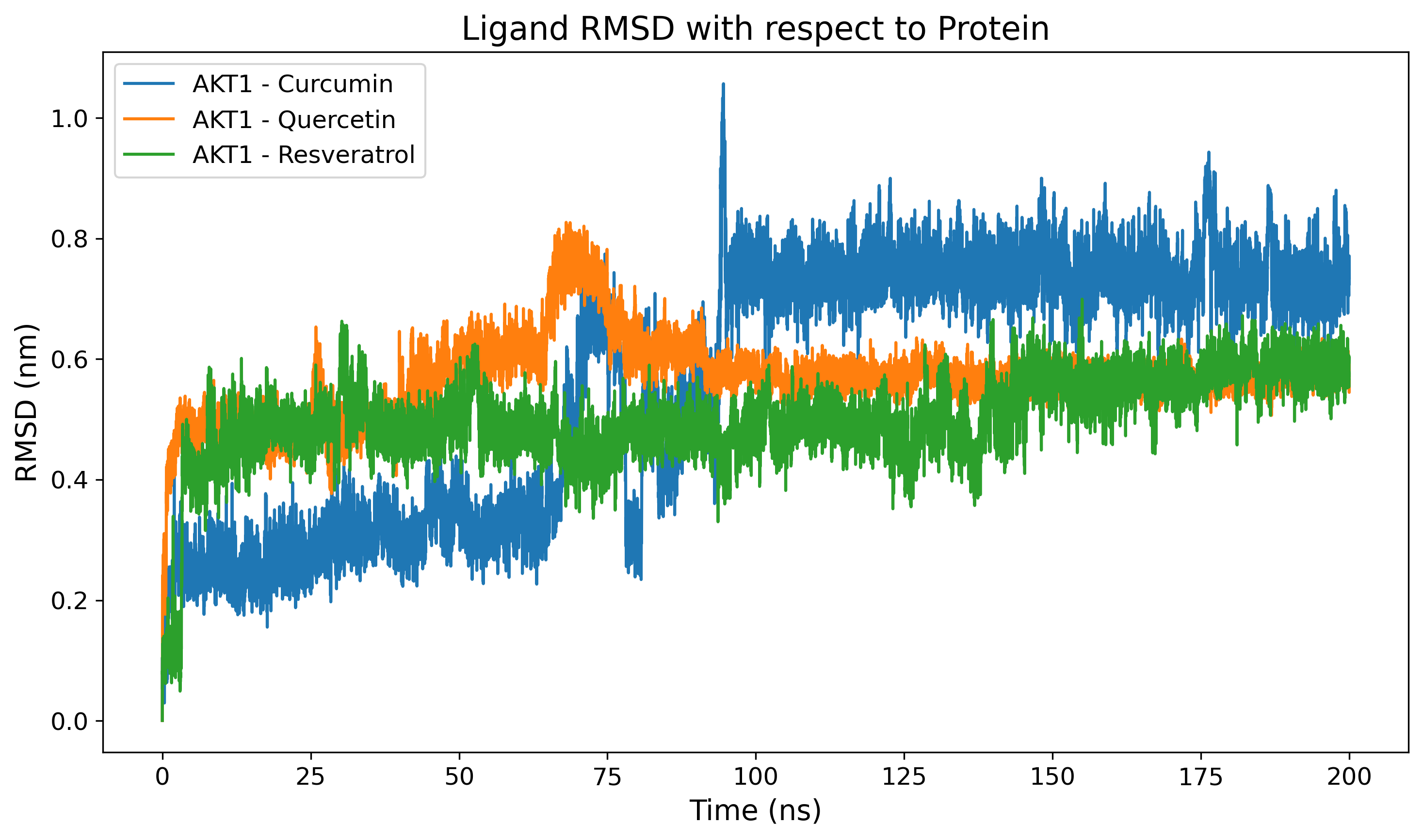
Ligand RMSD
Evaluates the positional stability of the ligand within the binding pocket.
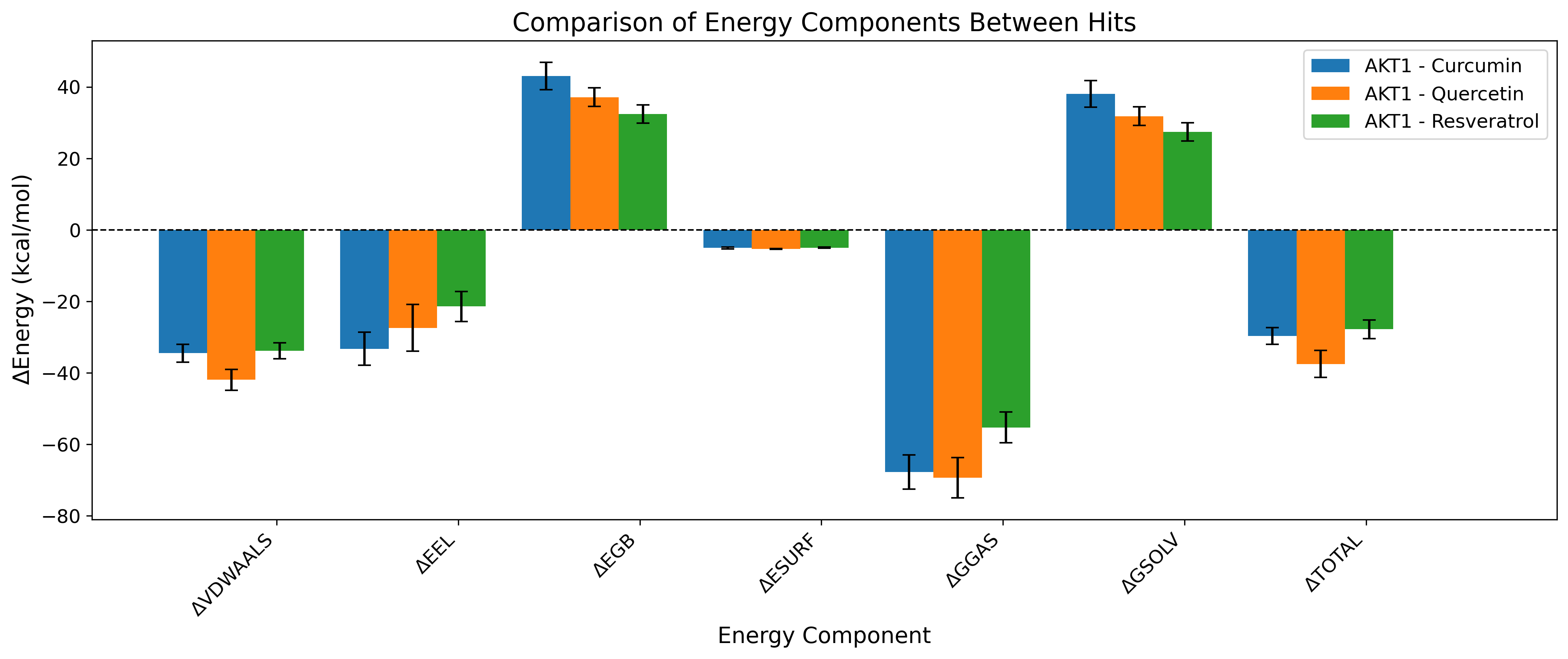
Binding Free Energy
Highly accurate MM/PBSA calculated binding affinities and energy decomposition.
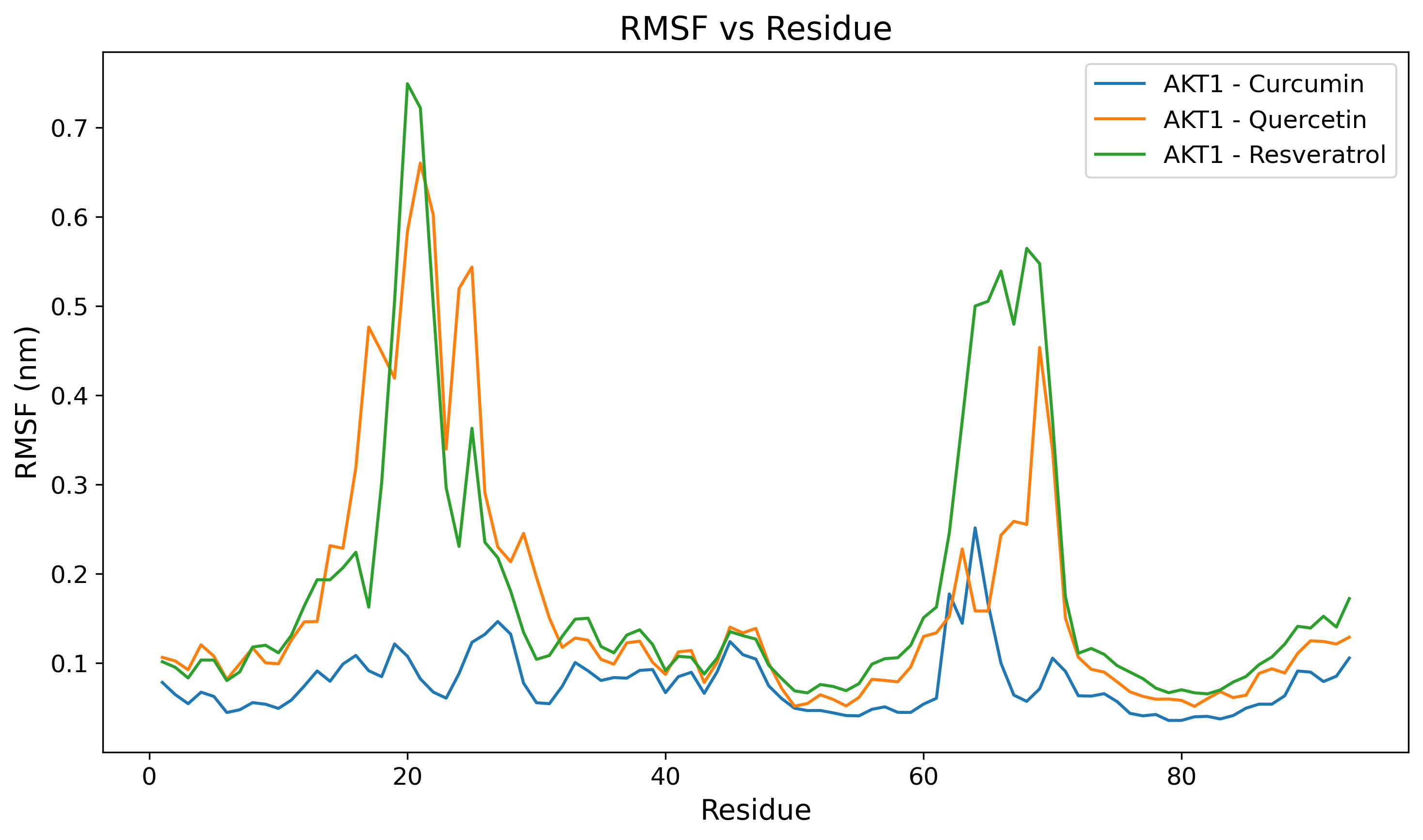
Protein RMSF
Root Mean Square Fluctuation detailing per-residue flexibility.
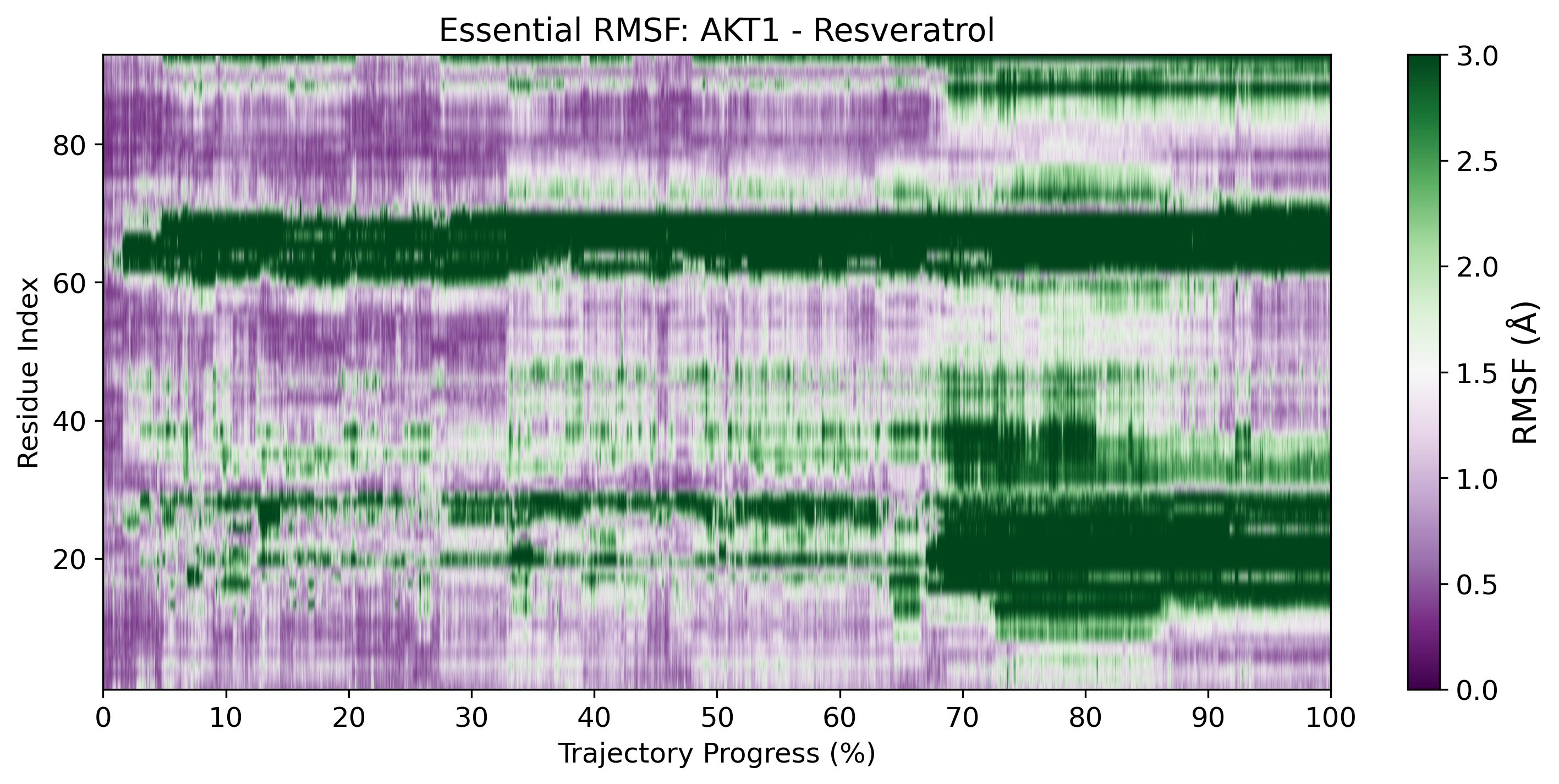
Essential RMSF (eRMSF)
Advanced fluctuation analysis isolating essential collective motions.
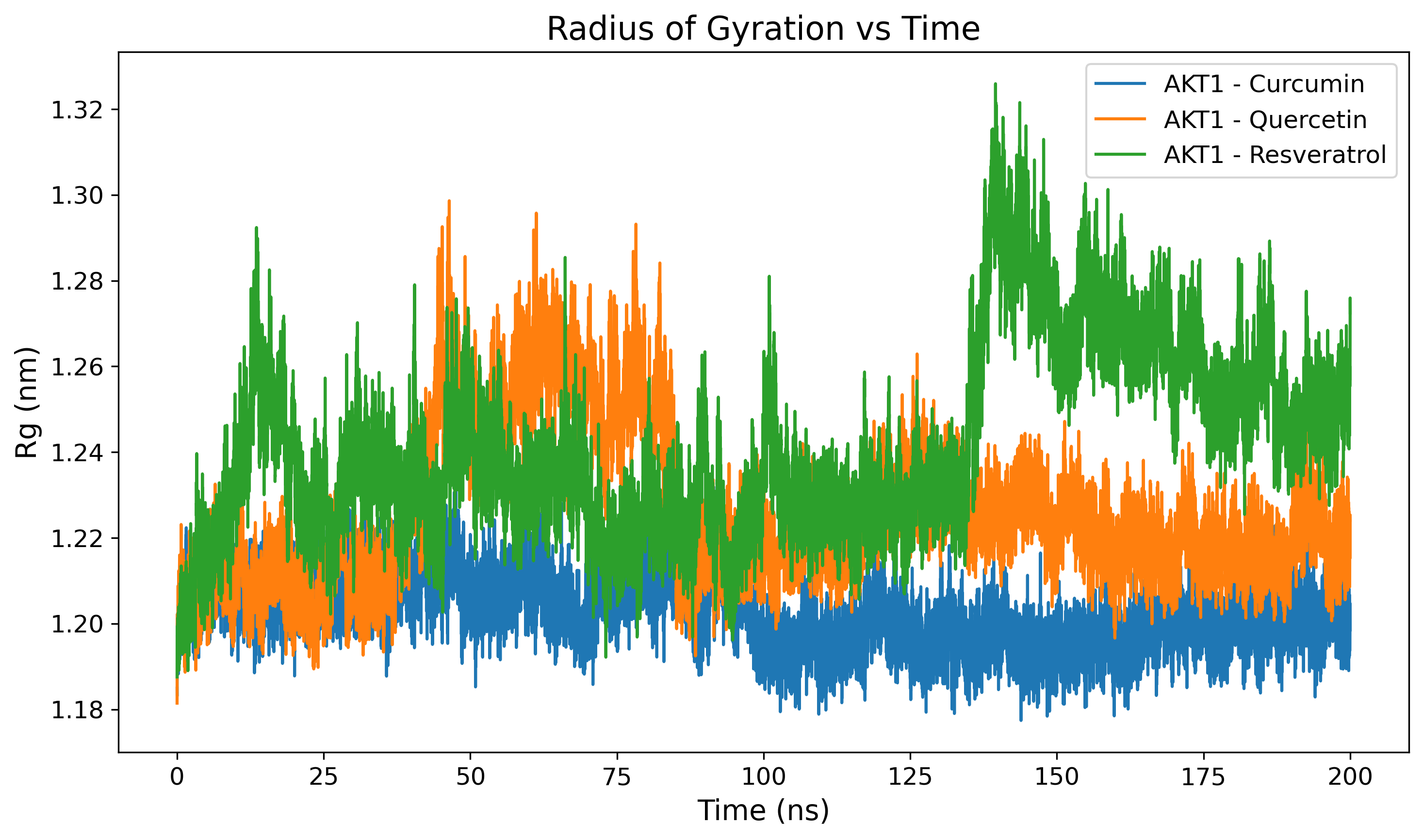
Radius of Gyration (Rg)
Measures the compactness and folding stability of the protein complex.
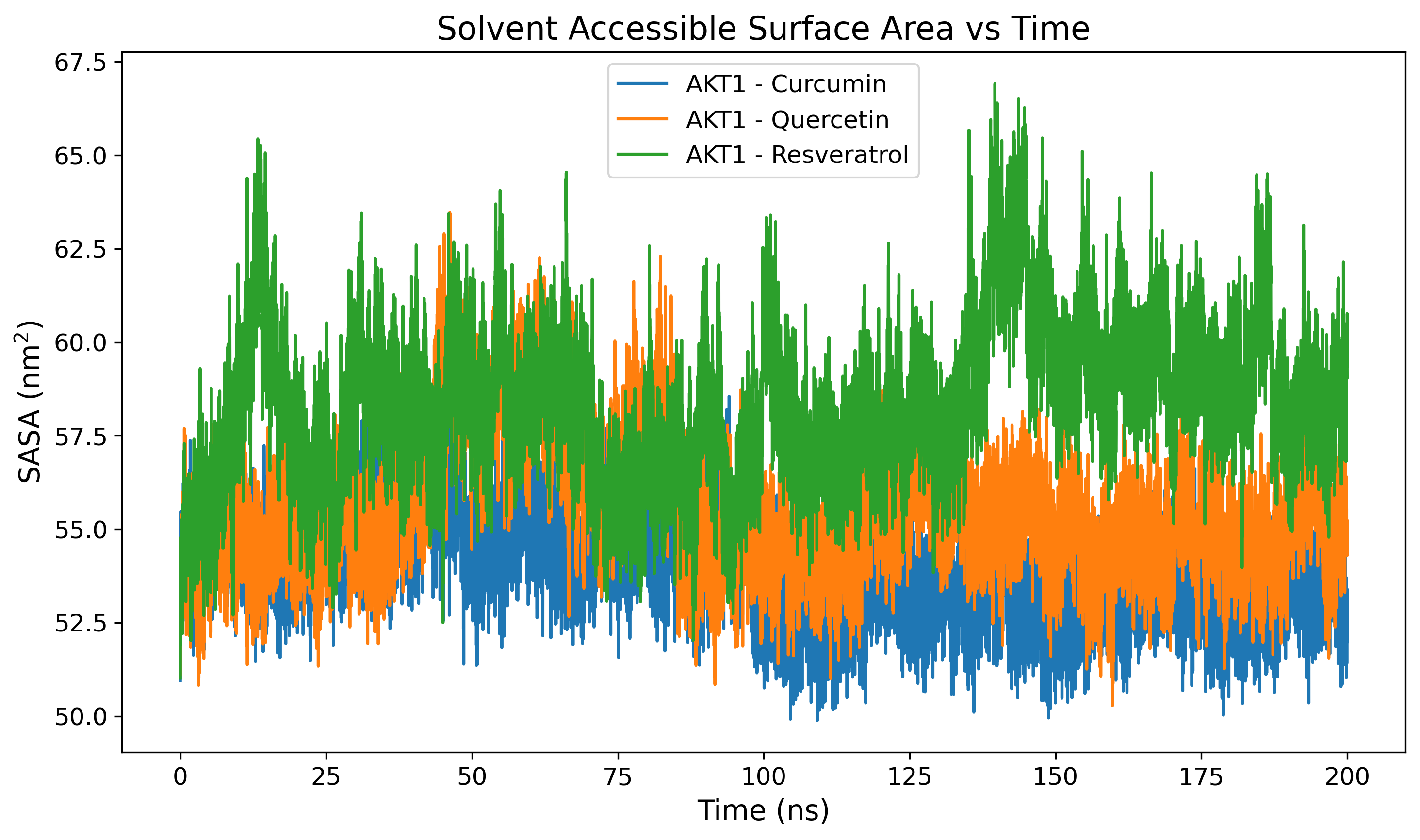
Solvent Accessible Surface Area
SASA analysis to monitor surface exposure and hydrophobic core integrity.
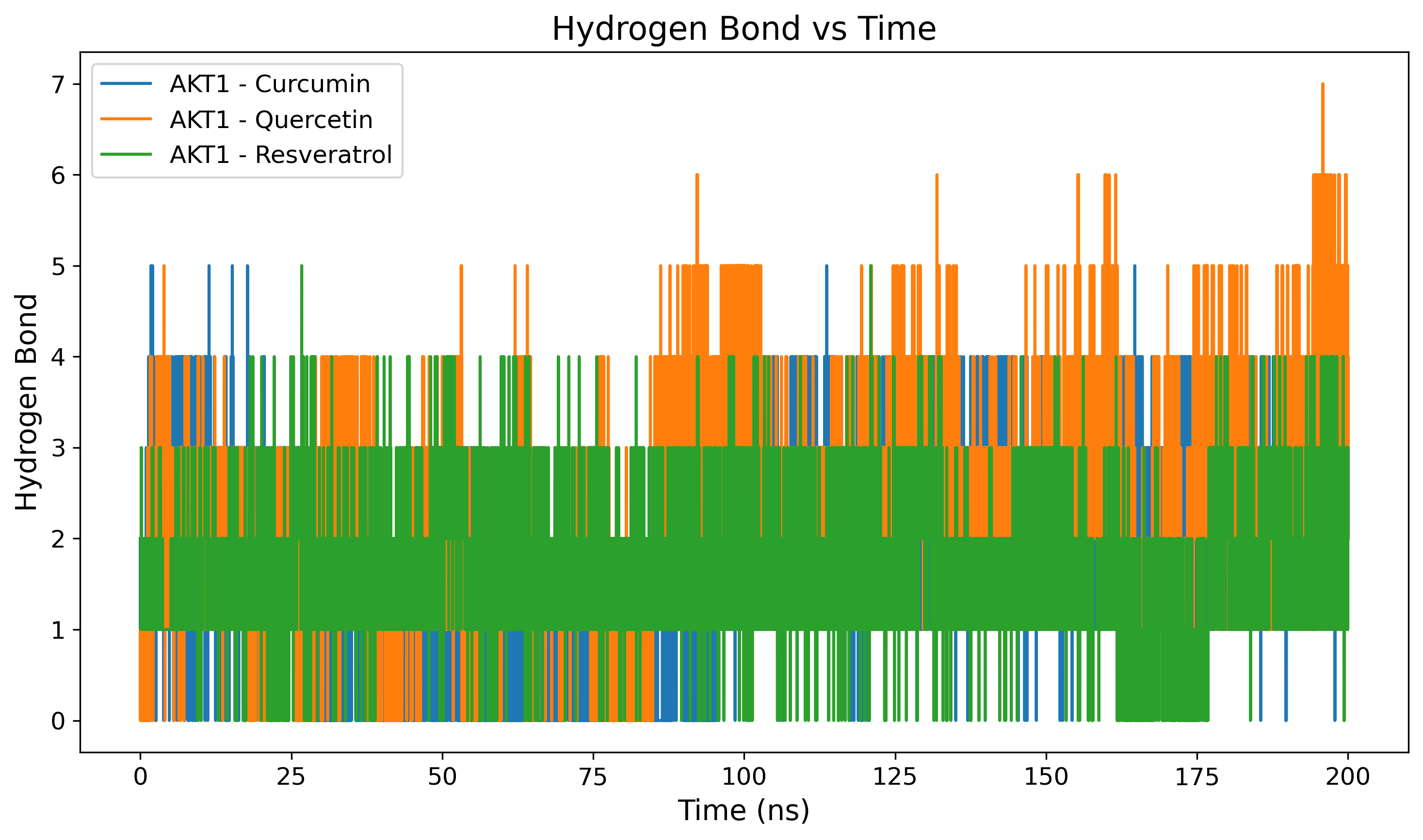
Hydrogen Bond Profiling
Tracks the number of intermolecular hydrogen bonds formed over time.
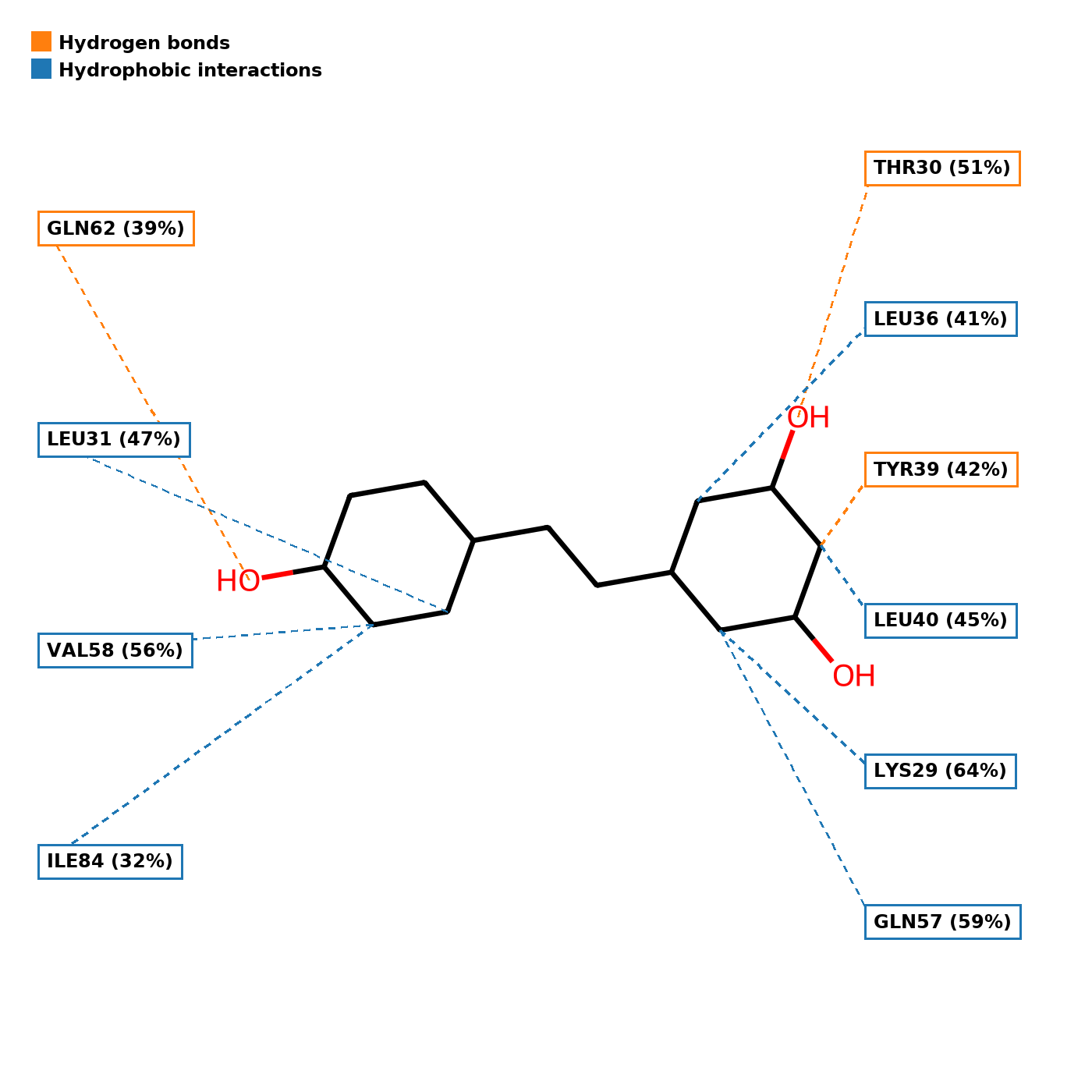
2D Interaction Plot
Schematic representation of persistent protein-ligand contacts.
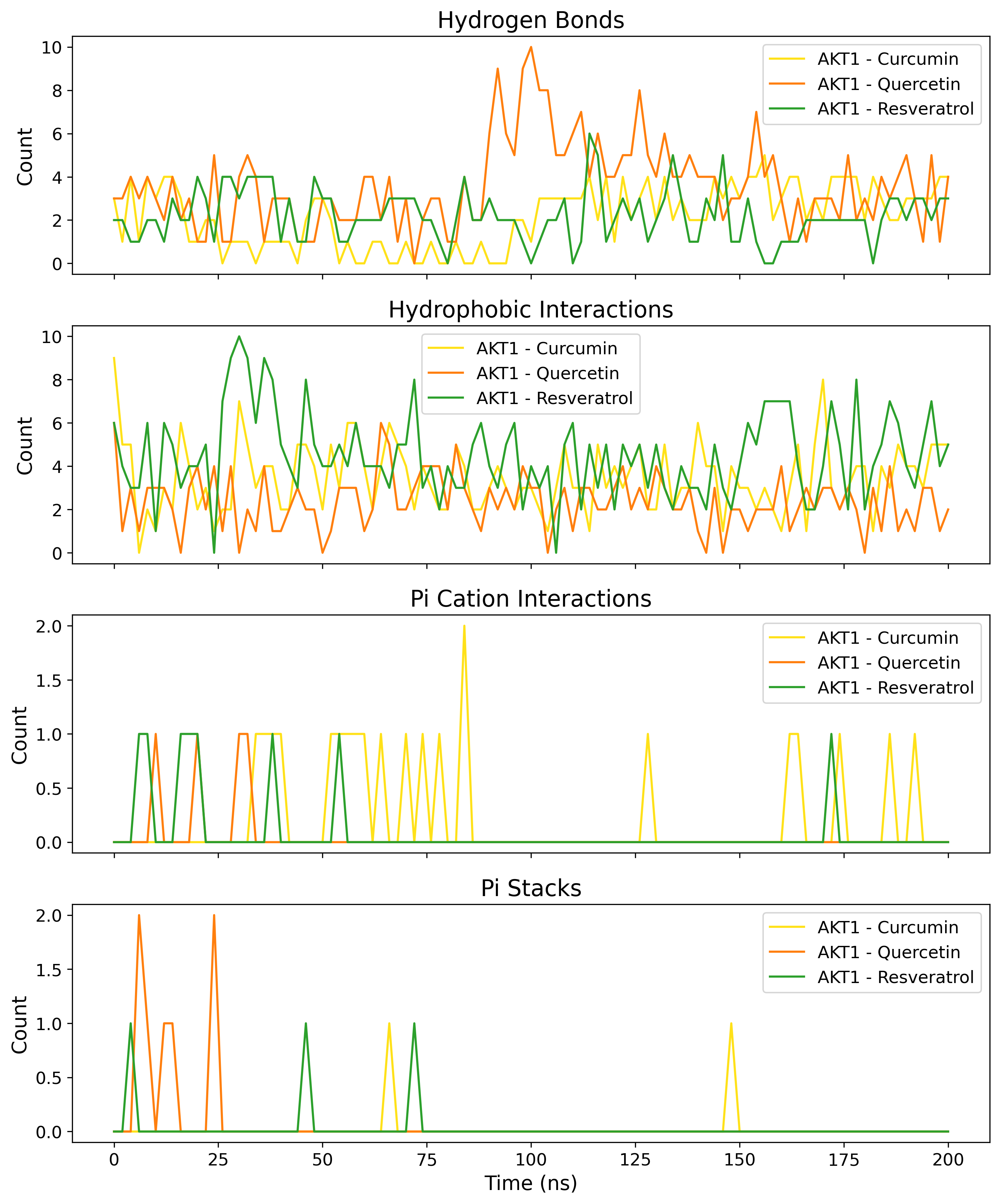
PLIP Profiling
Protein-Ligand Interaction Profiler identifying Pi-stacking and salt bridges.
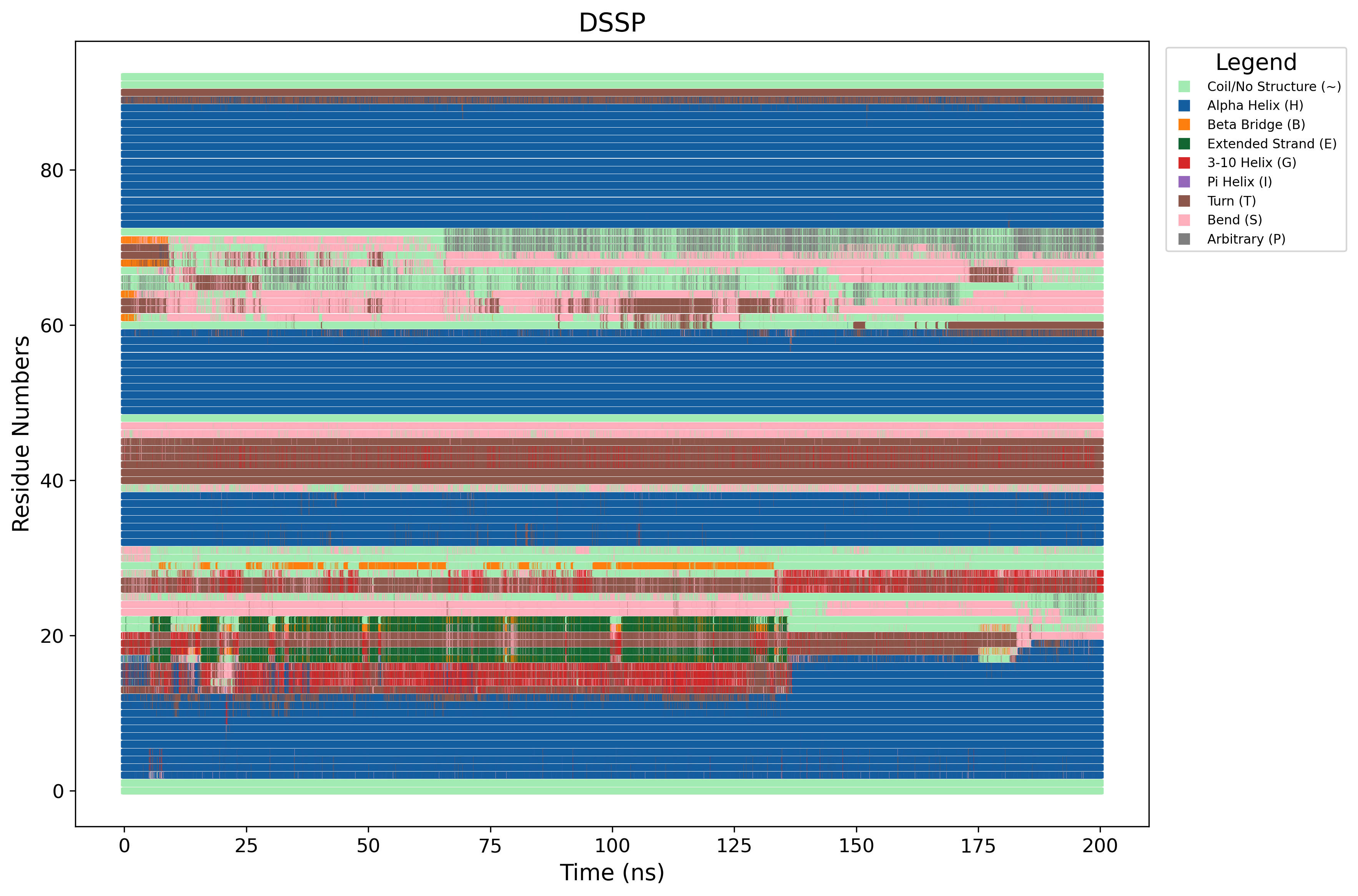
Secondary Structure (DSSP)
Time-evolution of alpha-helices and beta-sheets throughout the trajectory.
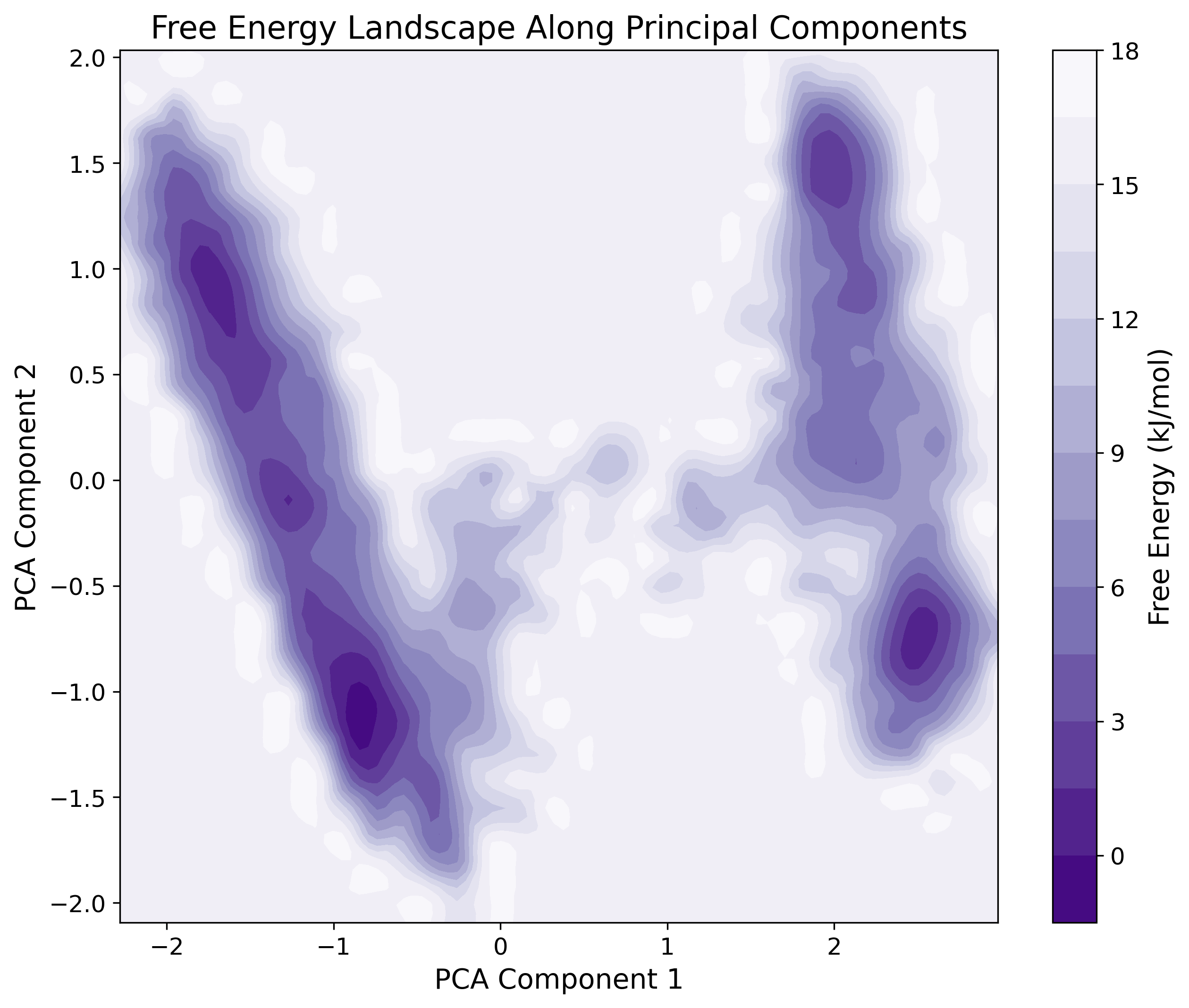
Principal Component Analysis
PCA identifying the dominant modes of structural variation.
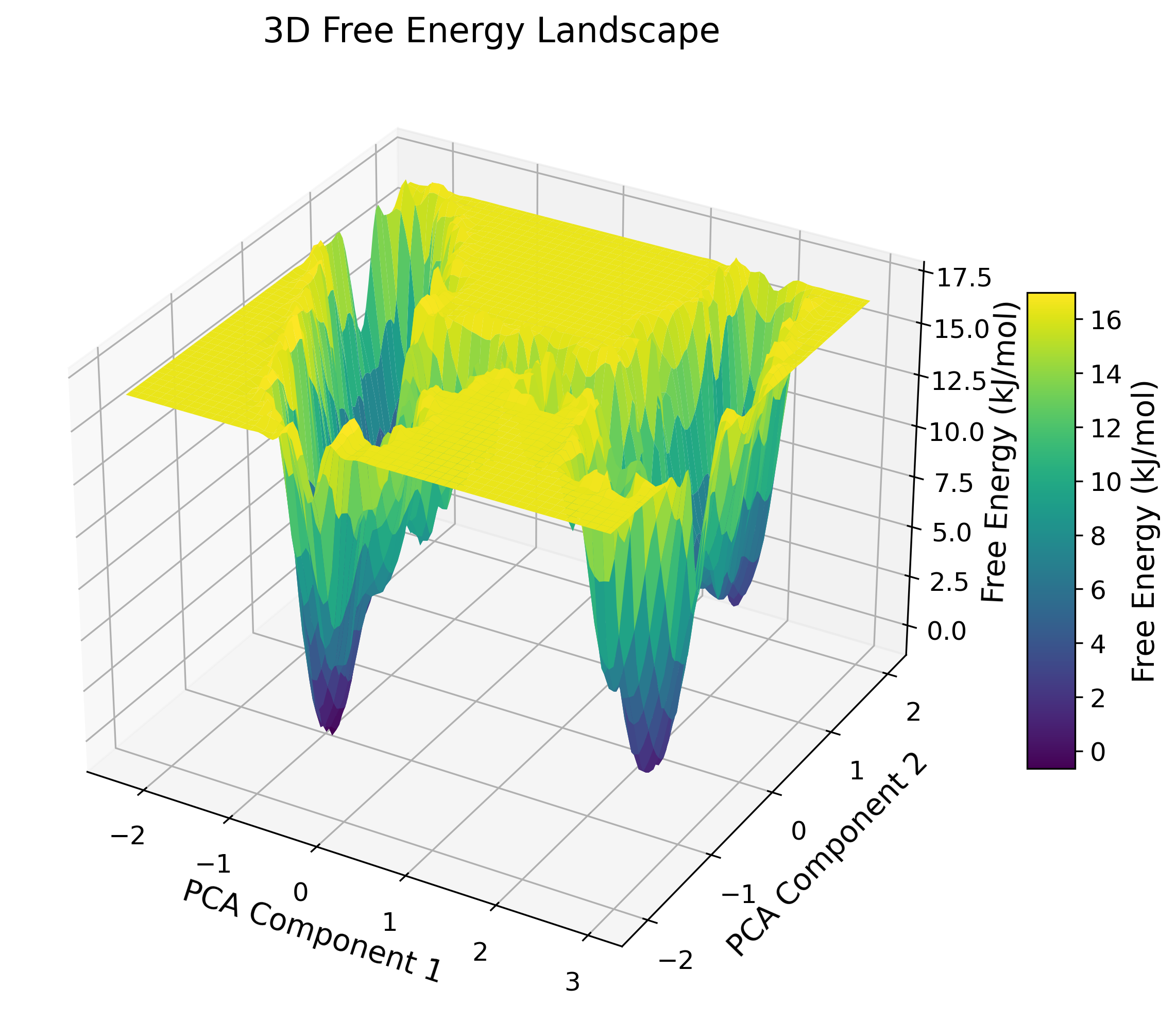
Free Energy Landscape (FEL)
3D contour mapping to identify metastable conformational states.
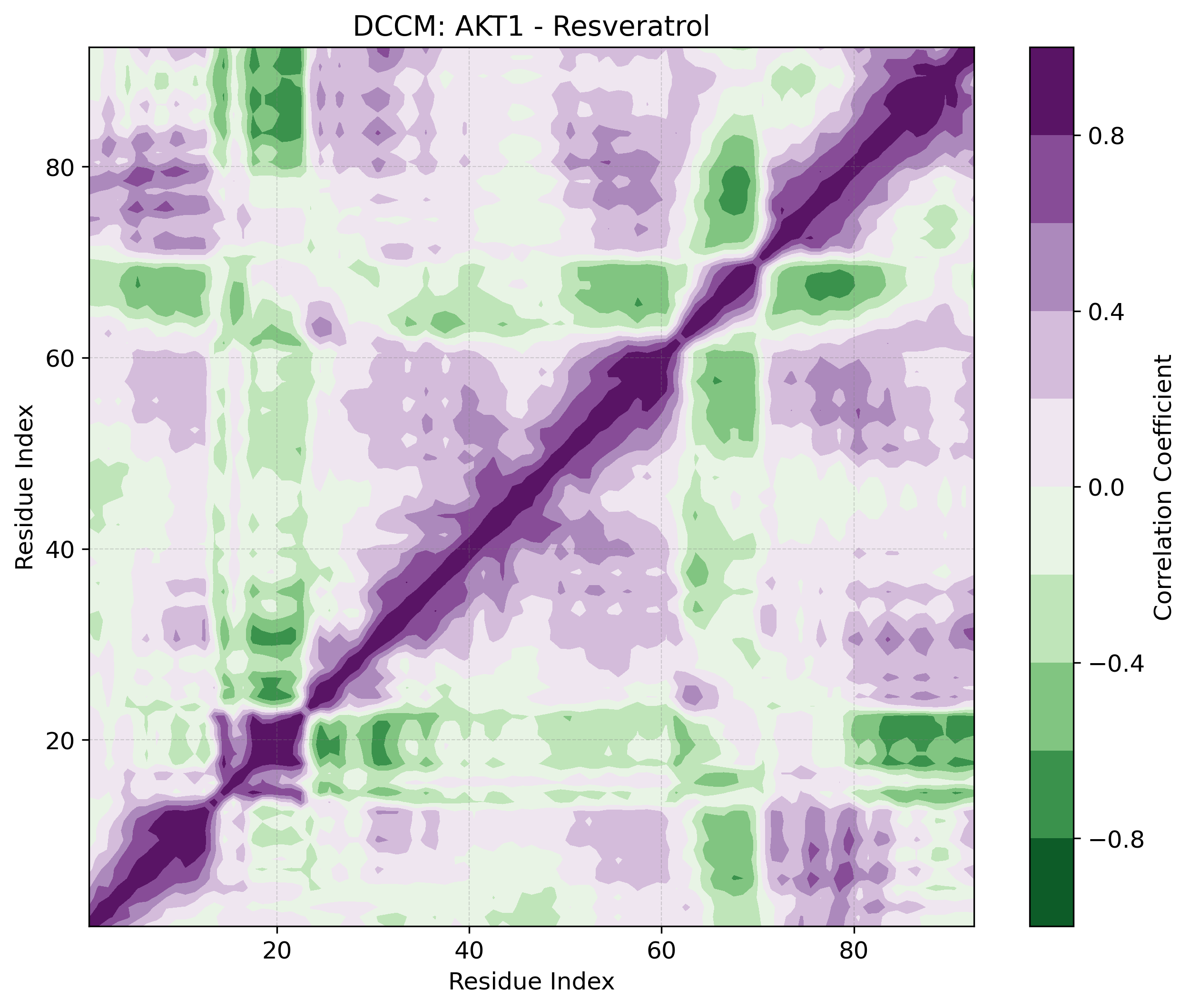
Dynamic Cross-Correlation
DCCM matrix revealing correlated and anti-correlated residue motions.
Statistics Summary
Comprehensive statistics summary (Mean and Standard Deviation) of each MD Parameters, like RMSD, RMSF, Rg, SASA, and more.
Loading table...
Input
Loading workflow configuration...
Pricing
Tools & Citations
| Tool Name | Purpose | Citation |
|---|